How to Build a PDF RAG Pipeline Without Text Extraction (Using Native PDF Embeddings)
Build a RAG pipeline that embeds raw PDF bytes with Gemini Embedding 2. No text extraction for embedding, vision-model OCR for the text you need, and better retrieval than extract-then-embed
Mar 18, 2026
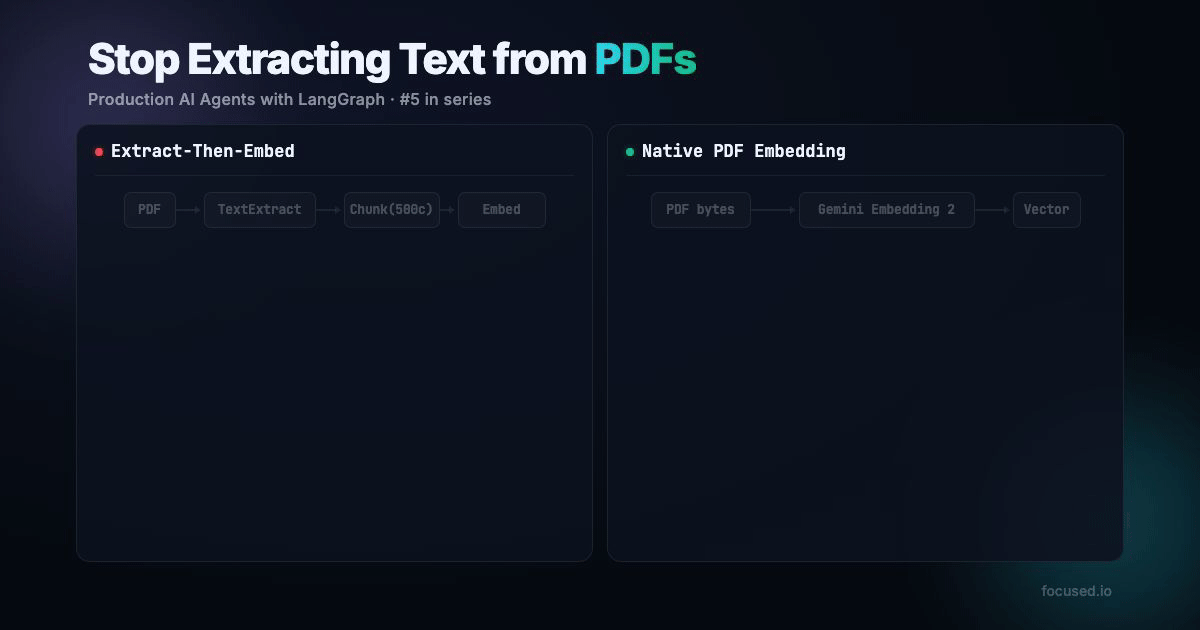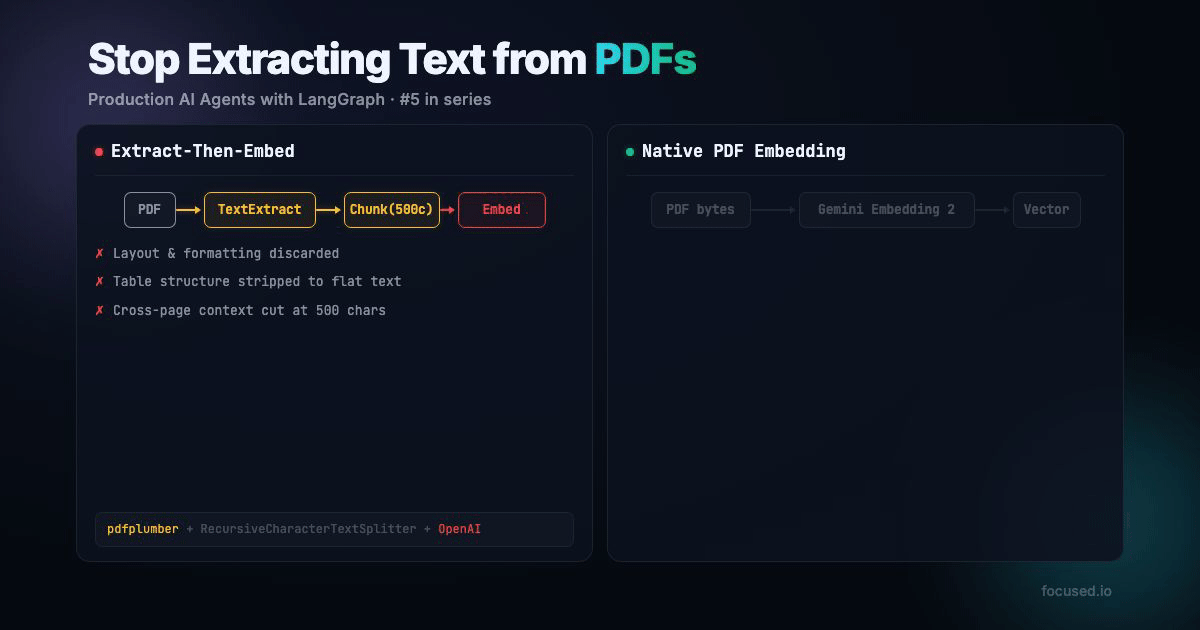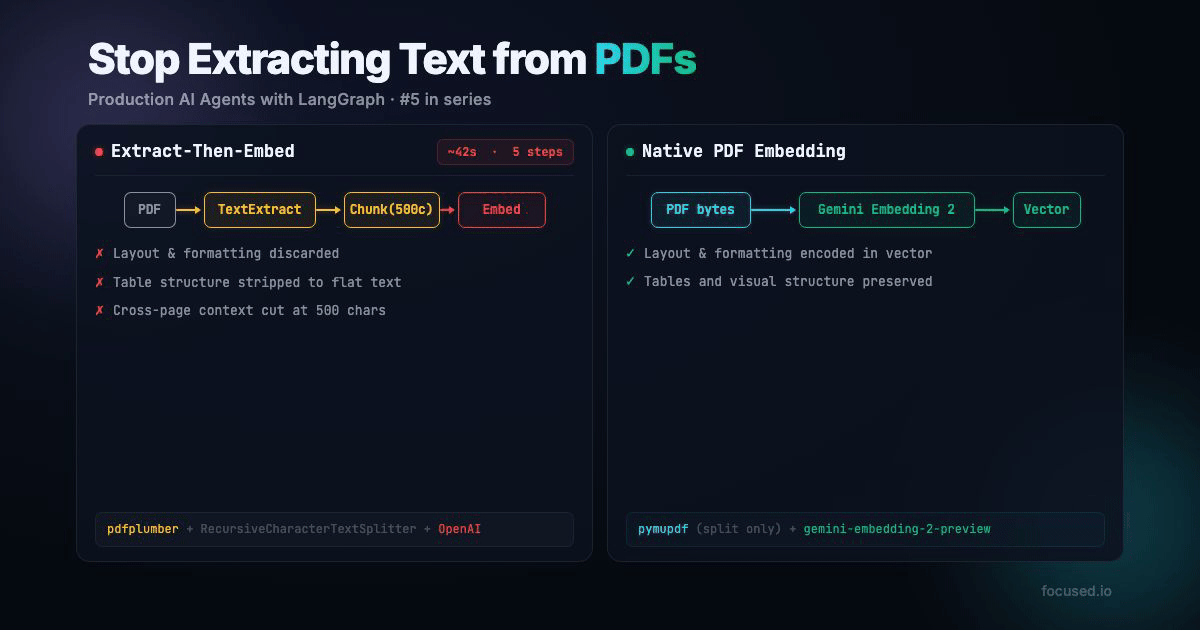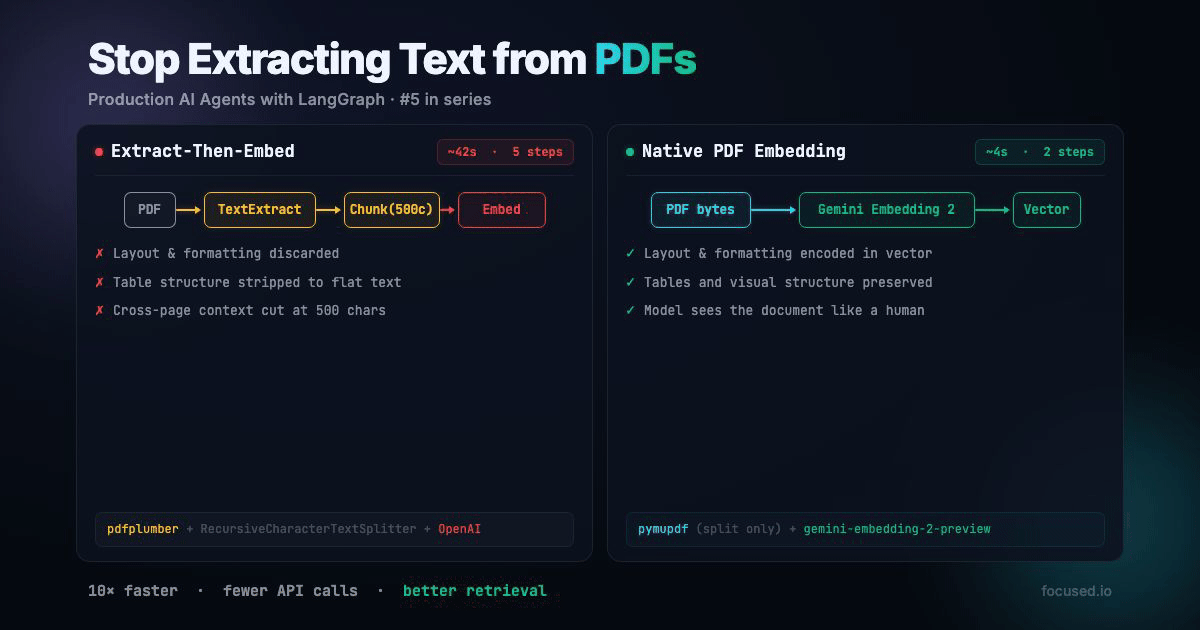
All RAG pipelines that handles PDFs does the same thing: extract text, chunk it, embed the chunks. The entire pipeline assumes that the text is the document. But text is often only a small part of the full story.
A court filing has headers, case numbers in specific positions, indented paragraphs that signal legal structure, signature blocks. A financial report has columns, footnotes, charts with captions. When you extract the text, you throw all that away. The embedding model sees a flat string. The document's visual structure, the thing people use to understand it, is stripped.
Google's Gemini Embedding 2 addresses this. You send it the raw PDF bytes. It reads the document just how a human would, layout, formatting, tables, everything, are all part of the embedding vector. Rather than text extraction and chunking. The embedding model sees the full document as you would.
We rebuilt a legal RAG pipeline around this. 215 scanned court documents, zero extractable text (every page is a scanned image), and the retrieval quality beats the extract-then-embed. This post shows how you can use LangGraph & Gemini Embedding 2 to build your own RAG.
The Ingestion Math
Traditional PDF ingestion has five lossy steps:
Native PDF ingestion has two steps:
That's faster, fewer API calls, no paid parsing service, and the embedding model sees more information. The main tradeoff is your minimum chunk granularity is 1 page instead of 500 characters. For documents with visual structure, which is most real-world documents, that's a better tradeoff.
The Architecture
PDF ──► pymupdf splits into ≤6-page groups
│
├──► Gemini Embedding 2 (raw PDF bytes → vector)
│ Sees layout, tables, formatting natively
│
└──► Gemini 2.5 Pro OCR (raw PDF bytes → text)
For full-text search + LLM context
│
▼
In-Memory Vector Store
(production: pgvector + tsvector)
│
User Query ──► Embed Query ──► Vector Search ──► Grade ──► Generate
▲ │
│ (not relevant)
└── Rewrite ◄────┘
The key insight: two models, same PDF bytes, different purposes. Gemini Embedding 2 produces the vector (captures visual semantics). Gemini 2.5 Pro produces the text (for keyword search and LLM context). Neither requires text extraction or image rendering and both accept raw PDF bytes.
State: Track Retrieval Quality
The state follows a corrective RAG pattern: retrieve, grade, rewrite if needed, and generate with citations.
from typing import TypedDict
from langchain_anthropic import ChatAnthropic
from langsmith import traceable
llm = ChatAnthropic(model="claude-sonnet-4-5-20250929", temperature=0)
class RAGState(TypedDict):
question: str
rewritten_query: str
documents: list[dict]
relevance_score: float
rewrite_count: int
max_rewrites: int
answer: str
citations: list[dict]
confidence: str
PDF Splitting: Pages as Natural Chunks
Gemini Embedding 2 accepts up to 6 PDF pages per request. For longer documents, split into page groups using pymupdf. This is the only thing pymupdf does, it's a splitter, not a text extractor.
from pathlib import Path
import fitz # pymupdf
MAX_PAGES_PER_CHUNK = 6
def split_pdf(filepath: Path) -> list[dict]:
"""Split a PDF into groups of up to 6 pages. Returns raw PDF bytes per group."""
doc = fitz.open(filepath)
chunks = []
for start in range(0, len(doc), MAX_PAGES_PER_CHUNK):
end = min(start + MAX_PAGES_PER_CHUNK, len(doc))
sub_doc = fitz.open()
sub_doc.insert_pdf(doc, from_page=start, to_page=end - 1)
chunks.append({
"pdf_bytes": sub_doc.tobytes(),
"page_numbers": list(range(start + 1, end + 1)),
"source": filepath.name,
})
sub_doc.close()
doc.close()
return chunks
A 20-page document becomes 4 chunks. A 3-page document is 1 chunk. Each chunk preserves cross-page context, a sentence spanning pages 5-6 is captured within a single embedding. Compare this to text chunking at 500 characters, where that sentence gets split at an arbitrary boundary.
Native PDF Embedding
This is the core innovation. Send raw PDF bytes to Gemini Embedding 2. The model natively understands document layout, headers, tables, formatting, visual hierarchy, and produces an embedding vector optimized for retrieval.
import math
from google import genai
from google.genai import types
gemini_client = genai.Client()
EMBEDDING_MODEL = "gemini-embedding-2-preview"
EMBEDDING_DIMS = 1536
def normalize(embedding: list[float]) -> list[float]:
"""L2-normalize an embedding vector (required for Gemini at non-default dims)."""
norm = math.sqrt(sum(x * x for x in embedding))
return [x / norm for x in embedding] if norm > 0 else embedding
@traceable(name="embed_pdf_chunk", run_type="embedding")
def embed_pdf_chunk(pdf_bytes: bytes) -> list[float]:
"""Embed raw PDF bytes using Gemini Embedding 2. No text extraction."""
result = gemini_client.models.embed_content(
model=EMBEDDING_MODEL,
contents=types.Content(
parts=[types.Part.from_bytes(data=pdf_bytes, mime_type="application/pdf")]
),
config=types.EmbedContentConfig(
task_type="RETRIEVAL_DOCUMENT",
output_dimensionality=EMBEDDING_DIMS,
),
)
return normalize(list(result.embeddings[0].values))
That's it. No RecursiveCharacterTextSplitter. No OpenAIEmbeddings. The PDF goes in, a vector comes out.
Asymmetric task types matter. Documents are embedded with RETRIEVAL_DOCUMENT, queries with RETRIEVAL_QUERY. Gemini Embedding 2 optimizes the embedding space differently for each, query vectors are tuned to find relevant documents, not to be similar to them. This is a meaningful retrieval quality improvement over symmetric embedding.
OCR_MODEL = "gemini-2.5-pro"
@traceable(name="ocr_pdf_chunk", run_type="chain")
def ocr_pdf_chunk(pdf_bytes: bytes) -> str:
"""OCR a PDF chunk by sending raw bytes to Gemini. No image rendering."""
response = gemini_client.models.generate_content(
model=OCR_MODEL,
contents=[
types.Content(parts=[
types.Part.from_text(
text=(
"Transcribe all visible text exactly as it appears. "
"Preserve paragraph structure, headings, and formatting. "
"Return ONLY the transcribed text, nothing else."
)
),
types.Part.from_bytes(data=pdf_bytes, mime_type="application/pdf"),
])
],
config=types.GenerateContentConfig(temperature=0, max_output_tokens=8192),
)
return response.text.strip()
No page.get_pixmap(). No base64.b64encode(). No rendering pages to PNG images. The model reads the PDF directly. This eliminates the memory overhead that causes OOM crashes on large documents, rendering a 67-page scanned PDF to 2x-resolution PNGs requires gigabytes of RAM. Sending the raw bytes requires kilobytes.
Ingestion: Two Models, Same Bytes
The complete ingestion function processes each PDF through both models in sequence:
DOCUMENT_STORE: list[dict] = [] # In-memory; use pgvector in production
@traceable(name="ingest_pdf", run_type="chain")
def ingest_pdf(filepath: Path) -> int:
"""Ingest a PDF: split pages, embed raw bytes, OCR for text."""
page_chunks = split_pdf(filepath)
for chunk in page_chunks:
embedding = embed_pdf_chunk(chunk["pdf_bytes"])
text = ocr_pdf_chunk(chunk["pdf_bytes"])
DOCUMENT_STORE.append({
"content": text,
"embedding": embedding,
"source": chunk["source"],
"page_numbers": chunk["page_numbers"],
})
return len(page_chunks)
Each chunk gets two things from the same PDF bytes: a vector that captures visual document semantics (for semantic search) and OCR'd text (for keyword search and LLM context).
Node 1: Retrieve
Vector search using cosine similarity on the native PDF embeddings. In production, this is a pgvector HNSW query. Here we use in-memory search to keep the example self-contained.
def cosine_similarity(a: list[float], b: list[float]) -> float:
dot = sum(x * y for x, y in zip(a, b))
norm_a = math.sqrt(sum(x * x for x in a))
norm_b = math.sqrt(sum(x * x for x in b))
return dot / (norm_a * norm_b) if norm_a > 0 and norm_b > 0 else 0.0
@traceable(name="retrieve", run_type="retriever")
def retrieve_node(state: RAGState) -> dict:
query = state.get("rewritten_query") or state["question"]
query_embedding = embed_query(query)
scored = []
for doc in DOCUMENT_STORE:
score = cosine_similarity(query_embedding, doc["embedding"])
scored.append({**doc, "score": score, "embedding": None})
scored.sort(key=lambda x: x["score"], reverse=True)
return {"documents": scored[:5]}
The retrieval here is identical to text-based RAG. The difference is what's in the vectors, PDF-native embeddings capture layout and visual structure, not only text semantics.
Node 2: Grade Relevance
Use structured output for clean boolean grading with with_structured_output.
from pydantic import BaseModel, Field
class RelevanceGrade(BaseModel):
"""Grade the relevance of retrieved documents to the user question."""
relevant: bool = Field(description="Are the documents relevant to answering the question?")
reasoning: str = Field(description="Brief explanation of the relevance assessment.")
grading_llm = llm.with_structured_output(RelevanceGrade)
@traceable(name="grade_relevance", run_type="chain")
def grade_node(state: RAGState) -> dict:
docs_text = "\n\n".join(
f"[Source: {d.get('source', 'unknown')}]\n{d['content'][:500]}"
for d in state["documents"]
)
grade = grading_llm.invoke([
{
"role": "system",
"content": (
"You are a relevance grader. Given a user question and retrieved documents, "
"determine if the documents contain information that can answer the question. "
"Be strict: tangentially related but non-answering documents are not relevant."
),
},
{
"role": "user",
"content": f"Question: {state['question']}\n\nRetrieved documents:\n{docs_text}",
},
])
return {"relevance_score": 1.0 if grade.relevant else 0.0}
Node 3: Rewrite Query
When retrieval fails, rewrite the query. Same pattern as text-based RAG — the rewriter doesn't know or care that the embeddings came from PDF bytes:
@traceable(name="rewrite_query", run_type="chain")
def rewrite_node(state: RAGState) -> dict:
response = llm.invoke([
{
"role": "system",
"content": (
"You are a query rewriter. The original query returned irrelevant documents. "
"Rewrite the query to be more specific and targeted. "
"Return ONLY the rewritten query, nothing else."
),
},
{
"role": "user",
"content": (
f"Original question: {state['question']}\n\n"
"Rewrite the query to find more relevant documents:"
),
},
])
return {
"rewritten_query": response.content,
"rewrite_count": state["rewrite_count"] + 1,
}
Node 4: Generate with Citations
The generator sees the OCR'd text — not the embeddings. Citations map to source filenames and page numbers:
@traceable(name="generate", run_type="chain")
def generate_node(state: RAGState) -> dict:
docs_text = "\n\n".join(
f"[{i + 1}] (Source: {d.get('source', 'unknown')}, Pages: {d.get('page_numbers', [])})\n"
f"{d['content']}"
for i, d in enumerate(state["documents"])
)
is_low_confidence = state.get("relevance_score", 0) < 1.0
system_prompt = (
"You are a document assistant. Answer the user's question using ONLY the provided "
"documents. Cite sources using [1], [2], etc. If the documents don't contain enough "
"information, say so — do not make up information."
)
if is_low_confidence:
system_prompt += (
"\n\nWARNING: Retrieved documents may not be fully relevant. "
"Be especially careful. Prefix with 'Note: Based on limited context.'"
)
response = llm.invoke([
{"role": "system", "content": system_prompt},
{"role": "user", "content": f"Question: {state['question']}\n\nDocuments:\n{docs_text}"},
])
citations = [
{"index": i + 1, "source": d.get("source", "unknown"), "pages": d.get("page_numbers", [])}
for i, d in enumerate(state["documents"])
]
return {
"answer": response.content,
"citations": citations,
"confidence": "low" if is_low_confidence else "high",
}
Graph Assembly
The graph follows the corrective RAG pattern. The only difference from text-based RAG is what's in the documents, PDF-native embeddings and OCR'd text instead of extracted-and-chunked text.
from langgraph.graph import END, START, StateGraph
from langgraph.types import RetryPolicy
def route_after_grading(state: RAGState) -> str:
if state.get("relevance_score", 0) >= 1.0:
return "generate"
if state["rewrite_count"] >= state["max_rewrites"]:
return "generate"
return "rewrite"
builder = StateGraph(RAGState)
builder.add_node("retrieve", retrieve_node, retry=RetryPolicy(max_attempts=3))
builder.add_node("grade", grade_node, retry=RetryPolicy(max_attempts=3))
builder.add_node("rewrite", rewrite_node, retry=RetryPolicy(max_attempts=3))
builder.add_node("generate", generate_node, retry=RetryPolicy(max_attempts=3))
builder.add_edge(START, "retrieve")
builder.add_edge("retrieve", "grade")
builder.add_conditional_edges(
"grade",
route_after_grading,
{"generate": "generate", "rewrite": "rewrite"},
)
builder.add_edge("rewrite", "retrieve")
builder.add_edge("generate", END)
graph = builder.compile()
Running It
if __name__ == "__main__":
from langsmith import tracing_context
# Ingest PDFs
ingest_pdf(Path("docs/sample.pdf"))
with tracing_context(
metadata={"pipeline": "native-pdf-rag", "version": "v1"},
tags=["production", "native-pdf-rag"],
):
result = graph.invoke({
"question": "What is this document about?",
"rewritten_query": "",
"documents": [],
"relevance_score": 0.0,
"rewrite_count": 0,
"max_rewrites": 2,
"answer": "",
"citations": [],
"confidence": "",
})
print(f"Answer: {result['answer']}")
print(f"Confidence: {result['confidence']}")
print(f"Rewrites: {result['rewrite_count']}")
Production Failures
1. 6-page limit truncation. A 67-page legal brief gets split into 12 chunks. If the answer spans chunks 3 and 7, neither chunk alone answers the question, and the grader marks both as irrelevant. Fix: after retrieval, expand top-scoring chunks to include sibling chunks from the same document. With a 200K context window, you can fit 3-4 complete documents.
2. OCR hallucination on degraded scans. Vision-model OCR is probabilistic. On low-quality scans, the model might "smooth over" illegible text instead of marking it uncertain. For legal or financial documents, this is dangerous. Fix: use the best available OCR model (Gemini 2.5 Pro, not Flash). Instruct it to output [illegible] rather than guessing. Verify critical fields (dates, amounts, case numbers) against structured metadata.
3. Embedding space mismatch between PDF and text queries. Document embeddings come from raw PDF bytes. Query embeddings come from text strings. Gemini's RETRIEVAL_DOCUMENT / RETRIEVAL_QUERY task types handle this asymmetry, but edge cases exist where a text query doesn't match the visual embedding well. Fix: hybrid search. Combine vector search (PDF embeddings) with full-text search (OCR'd text via tsvector/BM25) using Reciprocal Rank Fusion.
4. Memory pressure from sub-PDF creation. pymupdf's insert_pdf creates in-memory copies. For a 200MB scanned PDF, splitting into 6-page groups temporarily holds the full document plus all sub-documents in memory. Fix: process the sub-PDFs as they're created, then discard. Don't hold all chunks in memory simultaneously.
5. Rate limiting on Gemini embedding API. Each 6-page group requires one embedding call. A 215-document corpus with an average of 8 pages per document generates ~350 embedding calls. At Gemini's default rate limits, this can take 30+ minutes. Fix: batch processing with exponential backoff. Use the Gemini Batch API at 50% of the default embedding price.
Observability
Every function has @traceable with appropriate run_type. In LangSmith, a single query trace shows:
invoke_agent (chain)
├── embed_query (embedding) ← Query → Gemini Embedding 2
├── retrieve (retriever) ← Cosine similarity search
├── grade_relevance (chain) ← Structured output grading
├── generate (chain) ← Claude generates answerAnd an ingestion trace shows:
ingest_pdf (chain)
├── embed_pdf_chunk (embedding) ← PDF bytes → Gemini Embedding 2
├── ocr_pdf_chunk (chain) ← PDF bytes → Gemini 2.5 Pro
├── embed_pdf_chunk (embedding) ← next chunk...
└── ocr_pdf_chunk (chain)
The embedding run type is critical — it lets LangSmith show you embedding latency separately from LLM latency, so you can diagnose whether retrieval or generation is the bottleneck.
Evals
The eval suite tests three axes: does the retriever find content (retrieval quality), does the answer use only that content (faithfulness), and does the answer address the question (relevance).
from langsmith import Client, evaluate
from openevals.llm import create_llm_as_judge
ls_client = Client()
DATASET_NAME = "native-pdf-rag-evals"
if not ls_client.has_dataset(dataset_name=DATASET_NAME):
dataset = ls_client.create_dataset(
dataset_name=DATASET_NAME,
description="Native PDF RAG pipeline evaluation dataset",
)
ls_client.create_examples(
dataset_id=dataset.id,
inputs=[
{"question": "What is this document about?"},
{"question": "Who are the parties or people mentioned?"},
{"question": "What dates are referenced in the document?"},
{"question": "What is the main argument or conclusion?"},
],
outputs=[
{"expected_type": "summary"},
{"expected_type": "entity_extraction"},
{"expected_type": "date_extraction"},
{"expected_type": "argument_analysis"},
],
)
LLM-as-judge for faithfulness and relevance:
FAITHFULNESS_PROMPT = """\
Question: {inputs[question]}
Retrieved documents and generated answer: {outputs[answer]}
Rate 0.0-1.0 on faithfulness: Is every claim in the answer supported by the
retrieved documents? Unsupported inferences score 0.
Return ONLY: {{"score": <float>, "reasoning": "<explanation>"}}"""
faithfulness_judge = create_llm_as_judge(
prompt=FAITHFULNESS_PROMPT,
model="anthropic:claude-sonnet-4-5-20250929",
feedback_key="faithfulness",
continuous=True,
)
Custom deterministic evaluators:
def retrieval_has_content(inputs: dict, outputs: dict, reference_outputs: dict) -> dict:
"""Did retrieval return documents with actual OCR'd text?"""
docs = outputs.get("documents", [])
has_content = any(len(d.get("content", "")) > 50 for d in docs)
return {"key": "retrieval_has_content", "score": 1.0 if has_content else 0.0}
def rewrite_efficiency(inputs: dict, outputs: dict, reference_outputs: dict) -> dict:
"""Fewer rewrites = better initial retrieval. PDF embeddings should need fewer."""
count = outputs.get("rewrite_count", 0)
score = 1.0 if count == 0 else (0.7 if count == 1 else 0.4)
return {"key": "rewrite_efficiency", "score": score}
Run the suite with python evals.py. Check LangSmith for score distributions. If rewrite_efficiency averages below 0.7, your PDF embeddings aren't matching query semantics well — consider adding hybrid search with tsvector on the OCR'd text.
When to Use This
Use native PDF embedding when:
- Documents are scanned images (no extractable text)
- Layout carries meaning (legal filings, financial reports, engineering specs)
- You want to eliminate text extraction as a failure point
- You need fewer pipeline steps and API dependencies
Skip it when:
- Documents are plain text (Markdown, .txt) with no visual structure
- You need sub-page retrieval granularity (minimum chunk is 1 page)
- You need to embed thousands of documents per second (PDF processing is slower than text embedding)
- Your documents exceed 6 pages and need fine-grained cross-page retrieval
What This Replaces
The Bottom Line
The extract-then-embed pipeline was built around the limitations of text-only embedding models. Gemini Embedding 2 removes that limitation. If your documents have visual structure — and most real-world documents do — embedding the document directly produces better vectors than embedding extracted text. The pipeline simplifies from five lossy steps to one, with fewer dependencies, fewer failure modes, and better retrieval.
Technical References:

